You’ve mapped your protein network and validated expression, but the real question remains: Are your proteins interacting, and how much of each protein is free or interacting in your cell - or tissue sample?
The Challenge: Limitations of Traditional Protein Interaction Assays
Proximity Ligation Assay (PLA) is a powerful method used to detect protein–protein interactions (PPIs) directly inside cells or tissues. It works by using pairs of antibodies that are linked to short DNA strands. When two proteins are close to each other (typically less than 40 nm apart), the DNA strands can join together and be amplified through a process called rolling circle amplification (RCA), producing a fluorescent signal. This allows researchers to see where proteins interact at the single-cell level with high precision.
However, traditional in situ PLA has an important limitation: it only detects proteins that are interacting - the “bound” or “complexed” forms and provides no information about the free, unbound proteins. Because of this, PLA data shows only part of the picture. We can see that an interaction happens, but not how much of the protein is involved compared to the total amount. Without knowing both, it’s difficult to tell whether an interaction is a major feature of the cell or a rare event.
In addition, many common interaction assays have complex workflows that require specialized equipment and technical expertise. Some also depend on engineered protein expression, which can slow down experiments and make the results less representative of natural biological conditions.
Altogether, these challenges mean that traditional PLA and similar methods provide incomplete information about how proteins behave and interact dynamically within cells.
The MolBoolean™ Solution: Beyond Proximity Ligation
MolBoolean™ is a next-generation in situ proximity technology developed by Atlas Antibodies. It enables simultaneous detection and quantification of both free and interacting protein fractions in cells and tissues delivering a truly comprehensive spatial and quantitative profile of protein interactions (figure 1).
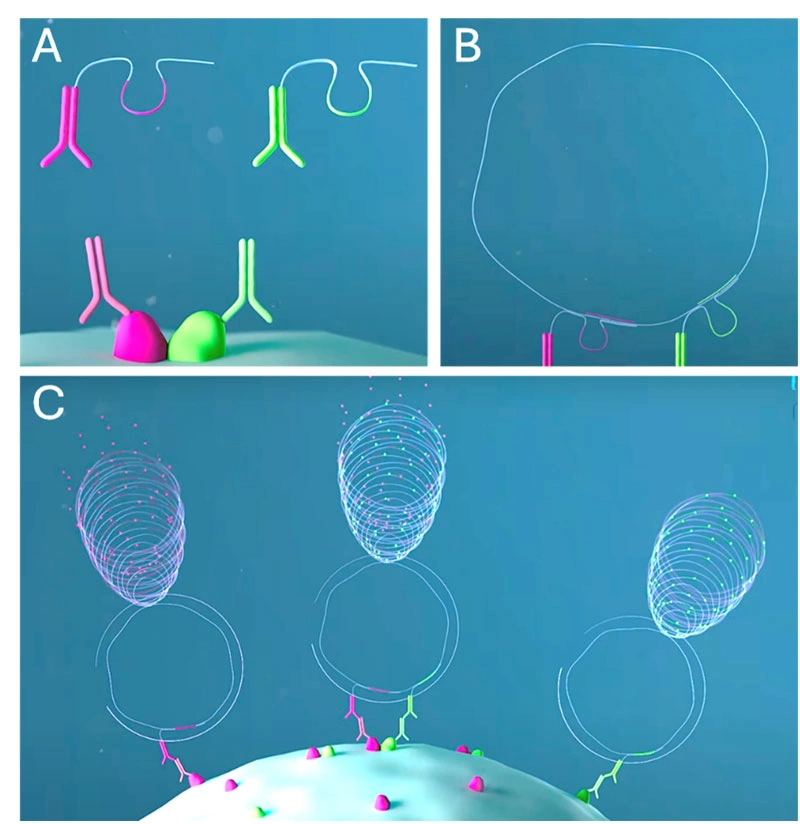
Figure 1.
A. Two primary antibodies enter the cell and bind to the proteins (magenta and green).
Two species-specific proximity probes, each conjugated with an information-containing DNA oligo, enter the cell and bind to the primary antibodies.
B. A pre-formed DNA circle (information receiver) hybridizes to the proximity probe oligos. The binding of the proximity probe oligos to the DNA circle generates double-stranded DNA Recognition sequences for a nicking enzyme which creates cuts in the DNA circle. Two identifier tag oligos get incorporated in the DNA circle at the cut sites by virtue of their complementarity to a loop-and hairpin region in the probe oligos.
C. The DNA circle templates are amplified through Rolling Circle Amplification (RCA forming long concatemeric Rolling Circle Products (RCPs). Depending on whether one or two fluorophore labeled tag-specific detection reporter oligos hybridize to the RCPs, three pools of proteins can be identified; two pools of un-bound proteins and the interacting, bound proteins.
MolBoolean™ solves the critical gap left by traditional assays by providing a full picture of protein interactions: free, bound, and total protein in one experiment which simplifies the workflow and eliminates the need for multiple assays.
How MolBoolean™ Transforms Your Workflow:
- Complete Data in One Assay: Quantify both free and interacting proteins in a single experiment. Normalize to total protein levels for reliable, biologically relevant data - supported by free image analysis tools and a simple step-by-step protocol.
- Universal Compatibility: Use your own primary antibodies (mouse and rabbit) or request custom secondary species for unique targets. No need for engineered proteins or specialized equipment.
- High Sensitivity: Rolling Circle Amplification (RCA) boosts fluorescence signal up to 1000-fold, enabling detection of low-abundance proteins.
- Consistent Results: Uniform molecular processing ensures reliable signal efficiency across experiments.
- Flexible Formats: Available as a ready-to-use kit and now also as a cost-effective Starter Kit which is ideal for evaluation and scaling up.
What This Means for Your Research
- Accelerate Discovery: Move from hypothesis to quantitative spatial data in a single, streamlined workflow.
- Expand Your Questions: Analyze both free and interacting protein pools critical for drug development, disease research, and systems biology.
- Adapt to Your Needs: MolBoolean™ is validated for both cells and tissue samples and can be tailored for your experimental requirements.
- Join a Growing Community: Researchers worldwide are leveraging MolBoolean™ for breakthroughs in neuroscience, oncology, immunology, and more.
To learn more about how you can apply the power of MolBoolean to your research, explore the technology on the Atlas Antibodies website or contact their technical support team.
-
What is MolBoolean™?
MolBoolean™ is a next-generation in situ proximity technology developed by Atlas Antibodies. It enables researchers to simultaneously detect and quantify both free and interacting protein fractions in cells and tissues, providing a comprehensive spatial and quantitative profile of protein interactions.
-
How does MolBoolean™ differ from traditional Proximity Ligation Assay (PLA)?
Traditional PLA detects only interacting (bound) proteins and does not provide information about free, unbound proteins. MolBoolean™ overcomes this limitation by quantifying both free and interacting proteins in a single experiment, giving a complete picture of protein dynamics.
-
What are the main advantages of MolBoolean™? The main advantages are:
- Complete Data: Quantifies both free and interacting proteins in one assay.
- High Sensitivity: Uses Rolling Circle Amplification (RCA) for a 1000-fold signal boost, detecting low-abundance proteins.
- Compatibility: Works with your existing primary mouse and rabbit antibodies, no specialized equipment needed.
-
Is MolBoolean™ compatible with different sample types?
Yes, MolBoolean™ is validated for both cell and tissue samples and can be tailored to specific experimental requirements.
